Thanks for checking out the paper, Max! I’m glad to hear that it prompted some ideas for your own research  . It’s very exciting to get feedback from other QS practitioners about how well techniques generalise across datasets, so thank you for all the details!
. It’s very exciting to get feedback from other QS practitioners about how well techniques generalise across datasets, so thank you for all the details!
When i read about previous day sleep impact, i’ve immediately did lasso on my data, sleep time from previous night survived and resulted in increase in adjusted-rsquared from 0.24 to 0.28.
Lagging features are surprisingly powerful, but (as I mention in the paper) there’s a lot of optimisation to be done in how to incorporate information from lags >1. But features with a lag of 1 are already just an easy way to boost predictive performance (and better understand the time series). I’m glad that worked for your model, but bear in mind that you’ve also added more parameters (lowering bias and increasing variance). Are those adjusted R^2 values on a validation set?
the idea of markov unfolding and overcoming missing data is pretty interesting, i’m going to try it when my noob skill in statistics will be improved…
Much of the complexity in my paper was because I was comparing these techniques to baseline, which requires some fiddly tricks to prevent data leaking. In terms of actually implementing them, it shouldn’t be too complex. Markov unfolding is a bit conceptually complex, but the actual implementation is pretty simple (in Python with Pandas), so long as your dataframe is already in the right format:
def markov_unfolding(df: pd.DataFrame, length=7):
""" Creates `length` new features for each column, corresponding to time lag.
"""
_df = df.copy() # Make a copy of original data
# Loop over numeric columns (except the target feature named 'target')
numeric_cols = _df.select_dtypes(np.number).columns
for c in [col for col in numeric_cols if 'target' not in col]:
# Apply shifts of up to `length` and stack them onto the new dataframe
for i in range(1, length+1):
_df[f'{c}_-{i}day'] = _df[c].shift(i)
return _df
(Based on the original implementation here.)
As for the missing data imputation, there are existing implementations in both R and Python for this. They handle most of the complexity for us  .
.
That looks weird for me. Bedtime isnt a measure of sleep quality - it’s something that affect sleep quality. Maybe it’s better to remove bedtime from sleep score (since you know the oura formula & weights, that’s shouldnt be a problem) and keep bedtime in dataset as a featuire which may influence sleep quality.
You’re definitely onto something here. I did consider extracting parts of the Oura score to construct a more tailored target feature. I decided against it because I wanted to keep the approach general (so that other Oura users could replicate my methods more directly). It would be interesting to look at how much noise vs. signal each component of the Oura sleep score contributes — perhaps I’ll tackle this one day in the follow-up work!
Keeping bedtime in a final feature (sleep score) have another problem - since the bedtime sometimes defined by your decision and not by features in data set, sleep quality will be distorted by including bedtime.
Exactly! This speaks to one of the major challenges of observational QS studies: the feedback loops make it really hard to figure out candidates for causal relationships. There is also quite a bit of variation between individuals here, I think. For instance, I’ve always struggled with insomnia and having a consistent sleep schedule. For me, my bedtime seems out of my control, so it made more sense to treat it as (part of) the dependent variable. But for someone who can fall asleep easily at whatever time they intend do, it’s much more of an independent variable (and therefore a feature). These nuanced decisions are part of why designing QS studies is so very challenging and bespoke to each individual. But maybe that’s also part of the fun!
according to my data oura sleep score is very noisy variable.
Yes, I’d say this is expected. Sleep quality (even in the abstract sense) is quite noisy as there are so many factors that contribute to it and sleep is still so minimally understood. Then we add in the noise introduced by sensors and using model predictions as proxy measures, etc.
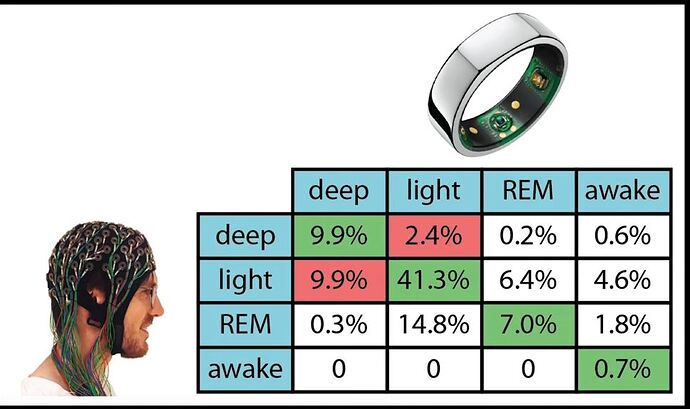
Sleep stages are pretty poorly detected
This seems to be consistent with the literature. Though I’m not sure that we even have good ways of measuring sleep stages in a lab — all our sensors are “guessing” in some sense. This obviously contributes to the high noise.
Total sleep time also poorly detected
This surprises me  My own experiences, comparisons by other QSers, and lab studies all find sleep duration (onset, interruptions, offset) to be quite accurate.
My own experiences, comparisons by other QSers, and lab studies all find sleep duration (onset, interruptions, offset) to be quite accurate.
Am I correct that you compared the Oura with a Dreem 2 headband? Both devices will have some noise, surely, but we can’t know which just from the Bland Altman.
So in a total - sleep stages, total sleep time and restlessness seems to have bad signal to noise ratio and i’m not sure in what degree oura sleep score represents sleep quality. There should be some useful signal, but data above raises a lot of questions.
Unfortunately, these things are all hard to measure (even with lab-grade polysomnography equipment), so we have to just deal with noisy variables here.
One workaround that might apply to you would be to build 2 identical explanatory models: one model using the Oura sleep score as a target, and one using the Dreem score as a target. That way, we know both are noisy, but any agreement between the feature importance/direction in the models is probably robust. I’d be very interested in seeing how consistent the models are, despite the targets not being in agreement.
Thanks for work you have done  i hope there will be more n=1 papers with deep dive like that
i hope there will be more n=1 papers with deep dive like that
Thanks for all the great feedback and for replicating some of the analysis!  I’m looking forward to seeing your future experiments!
I’m looking forward to seeing your future experiments!
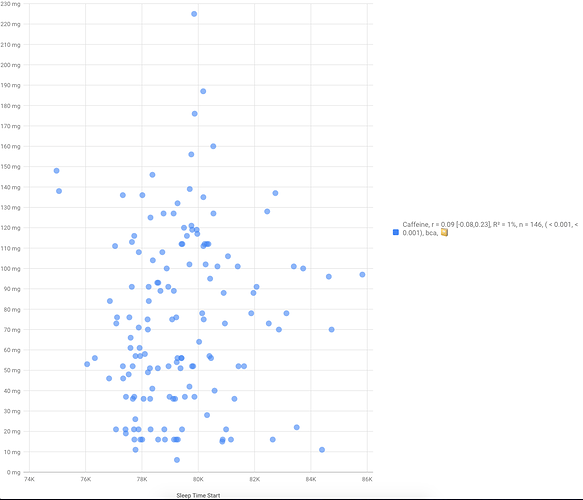
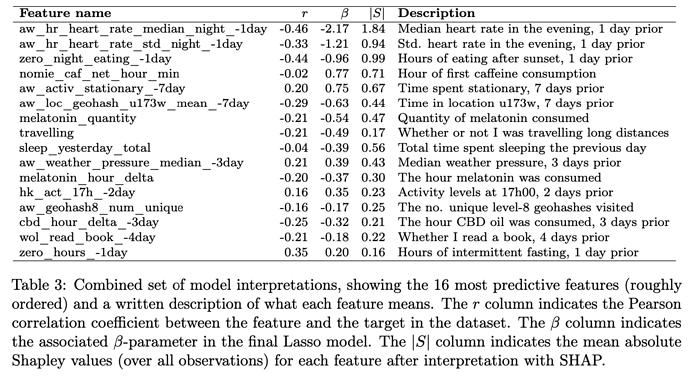

 Actually i did that in some degree, but not in a solid way like you
Actually i did that in some degree, but not in a solid way like you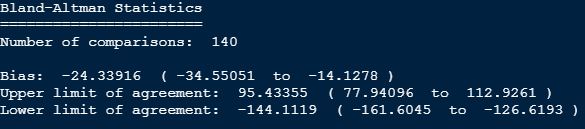

 My own experiences,
My own experiences,