I have results from sampling the same stool in 2 places with ubiome. I am happy to prepare results if folks are interested. Anything specific you would like to see – there is a lot of data?
Hi Isaac,
I am curious what differences you see in the two results. Were you surprised? How about compared to your full sample history spanning months.
Any idea what meal the samples were taken from (markers like beet coloring or undigested corn kernels)? Does your diet change significantly between meals?
Can you associate the samples with different Rx intake?
Was your health status stable during the time frame for digestion and sampling?
Do you have any information on the Measurement System Analysis (reliability and repeatability) of the sample analysis done in uBiome’s lab?
I assume both samples were taken at approximately the same time so that should not be a factor. (I recall that exposure to oxygen is problematic for some of the microbes). I also assume the sample storage tube has a preservative that fixes the relative counts so they don’t grow on the way to the lab, both samples arrived on the same day and were tested on the same equipment by the same person.
Ok, here comes a big dump of information (pun intended, har har har)…
Here’s what i found (and I am a little surprised):
- my ubiome wellness score was 88% for one sample and 96% for the other. In my mind, a big difference.
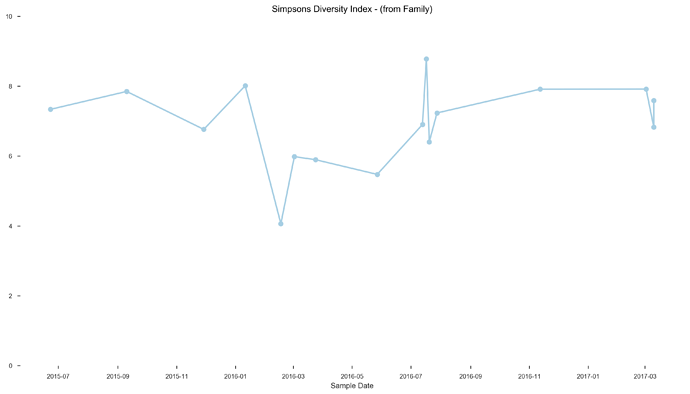
- ubiome diversity was 20th and 53rd percentiles. Also a big difference in my mind.
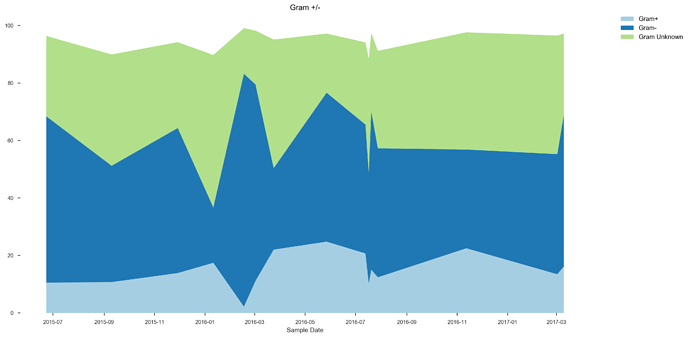
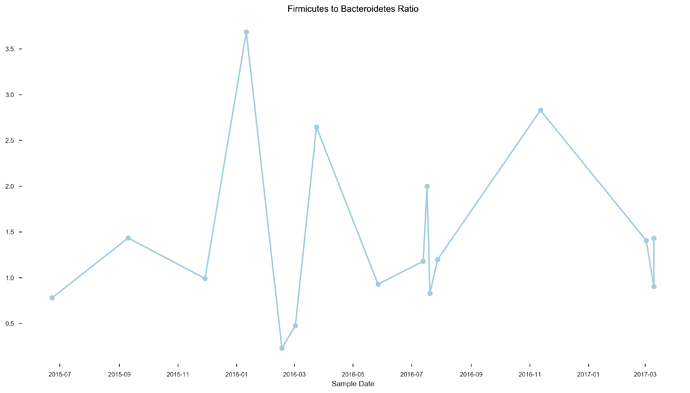
- F/M ratio was 0.9:1 and 1.4:1
- no akkermansia and very little probiotics detected in the one sample, the other sample had good amount of both.
How about compared to your full sample history spanning months.
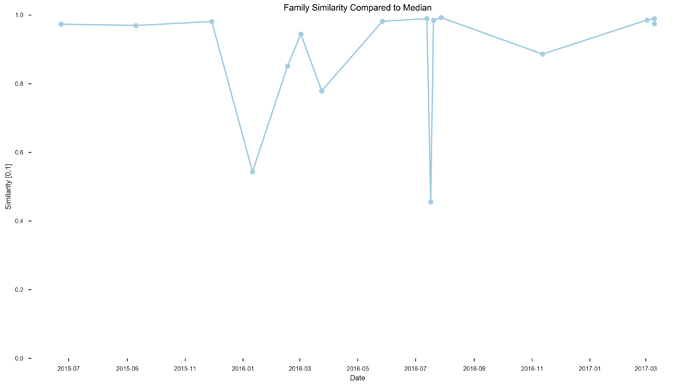
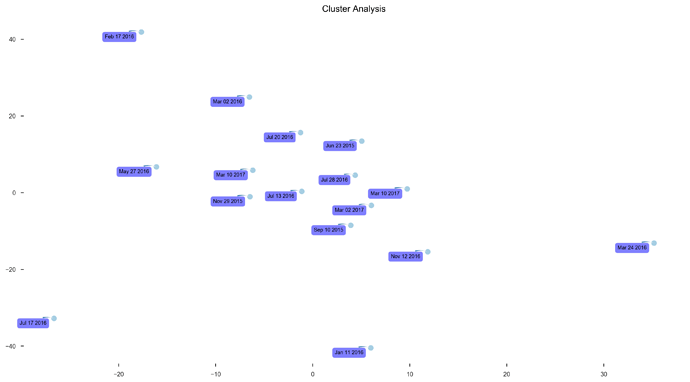
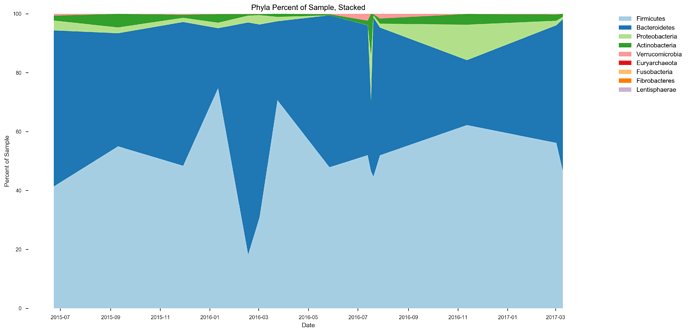
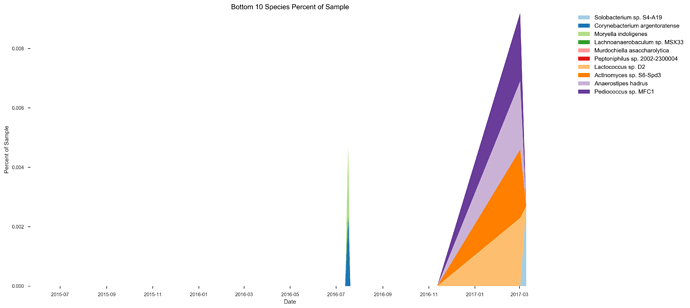
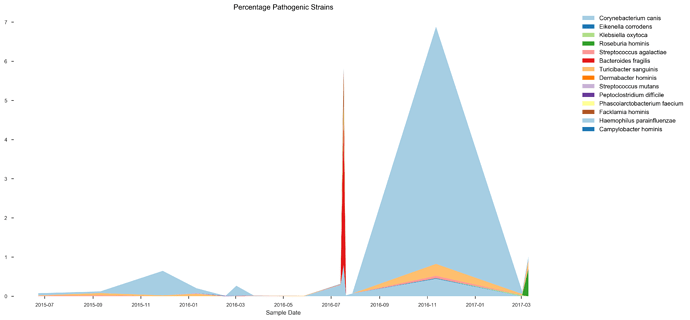
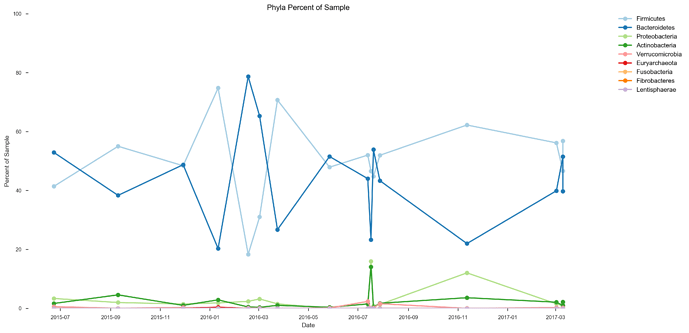
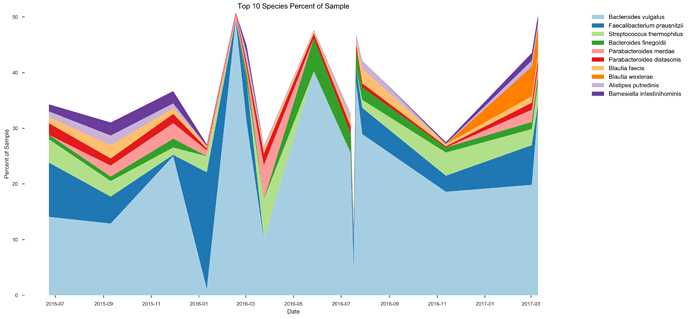
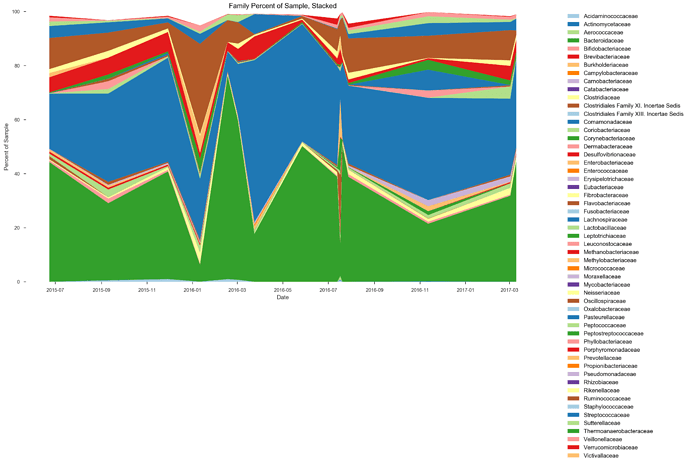
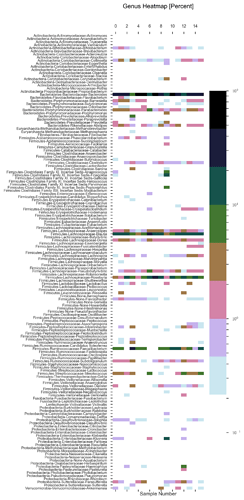
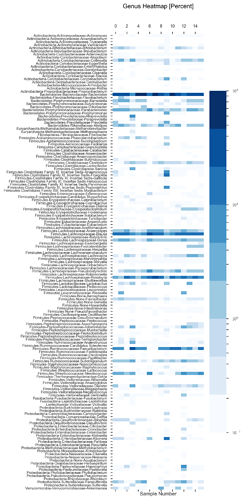
See plots here, specifically, the last 2 samples are from the same piece of stool sampled at 2 different sites, roughly first quarter and last quarter (also added inline to message): ubiome_longitudinal_analysis/sample_data at master · isaacgerg/ubiome_longitudinal_analysis · GitHub
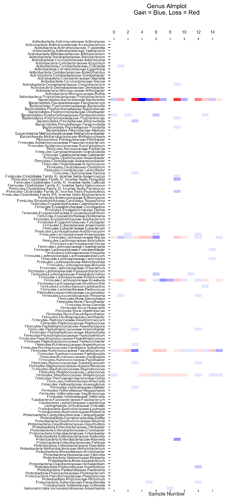
If you look at the genus Almplot and the genus heatmap of the last 2 samples, you can see the differences most clearly in my mind.
I just upgraded to matplotlib 2.0 so my stylessheet is messed up. Please bear with me (for example not grids on plots).
Any idea what meal the samples were taken from (markers like beet coloring or undigested corn kernels)?
No idea. Nothing stood out to me. I have pictures of the sampling process if are really interested so you can see for yourself.
Can you associate the samples with different Rx intake?
I am in the process of looking at this now. So far, the anova hasnt shown anything but i just got back 3 more samples. I likely wont have this analysis done until later summer since Im working on another paper at the moment.
Was your health status stable during the time frame for digestion and sampling?
Mild IBS
Do you have any information on the Measurement System Analysis (reliability and repeatability) of the sample analysis done in uBiome’s lab?
Nothing but anecdotes so I will not say anything. You can see the number of reads for each sample in the .json files. The first part of the stool had worse diversity score of the 2 (ref comments above) and the number of reads was:82321. the second part of the stools had 73785 reads. So both seem fine to me in terms of quality. In my plots, the first part of the stool occurs first in time of the 2 samples. (json files 101419 and 101960 respectively in my repo – they are also labeled in the json notes).
I assume both samples were taken at approximately the same time so that should not be a factor. (I recall that exposure to oxygen is problematic for some of the microbes). I also assume the sample storage tube has a preservative that fixes the relative counts so they don’t grow on the way to the lab, both samples arrived on the same day and were tested on the same equipment by the same person.
Both taken at the same time. My hypothesis is that the biome of the outer part of the stool differs significantly from the inner part of the stool. My next samples will specifically test this.
Hi Isaac,
I will pass on your offer of pictures of your sampling technique. It may be difficult to prevent surface contamination of the interior samples. Good luck on a clean catch.
That level of variation doesn’t seem surprising, but it does not bode well for the use of this tool to tune up your system. One would hope that while experiencing mild IBS, that you would not get a 96% wellness score on any measurement you were using to improve (IMHO). uBiome health may be more of a necessary but not sufficient type of condition. Maybe something to check if you get a flare up.
Have you considered other measurements?
That level of variation doesn’t seem surprising, but it does not bode well for the use of this tool to tune up your system.
I would disagree and I’ll offer this reason, I bet that when i get the smartgut report back from each of these samples (which will be a few more weeks), it will show signifiantly different results. This is based on looking a previous smartgut reports.
ne would hope that while experiencing mild IBS, that you would not get a 96% wellness score on any measurement you were using to improve (IMHO).
I’m not so sure that’s true. IBS is a problem with the small bowel that manifests as symptoms of the large bowel, mostly that gas is produced in the small bowel, migrates to the large bowel, and then interacts with the large bowel either speeding up or slowing down transit time.
Maybe something to check if you get a flare up.
I was having a flare at this time.
You have to keep in mind that IBS is poor layman medical term that in 20 years will likely no longer be used. “IBS” as we understand it now is actually a set of conditions all lumped under the same name. Its like saying someone has cancer but not specifying what kind – all cancers are not created equal.
Based on my testing and the many doctors I have talked to, I have visceral hypersensitivity. I have thought about getting antibody tests or doing a mannitol ingest test but am still reading up on the science to understand the sensitivity and specifity of these tests (i.e. are they useful for diagnosing). Its also worth mentioning that there are no good predictors of IBS right now from stool samples, but there are very good predictors from jejunal aspirates.
It looks like this post has received a lot of views over the past day. Is this an area people would be interested in having me host a youtube stream to answer questions and speak about my interpretation of the results?
I certainly would be.
I’m happy to set something up. Is there an easy way to poll people for availability?